Welcome to LocalMove webserver !
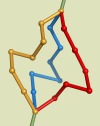 Local move is a Monte-Carlo approach to the problem of finding best-fitting lattice models for biopolymers.
Local move is a Monte-Carlo approach to the problem of finding best-fitting lattice models for biopolymers.
Starting from an initial discrete model (3D self-avoiding walk), it performs a sequence of elementary random
alterations, called Local Moves. The original atoms coordinate, input as a
PDB file, are then used to decide whether or not to accept the candidate
move, either deterministically in a greedy way or stochastically by performing a simulated annealing. Through iterating the
process for a reasonable amount of time, a model is obtained whose inter-atom distance is very close to the original
model.
 Using this this webserver, it is possible to:
Using this this webserver, it is possible to:
Run a LocalMove experiment
Monitor fitting progress in real time (Requires Java)
Retrieve the best-on lattice output
Additionally, you can reproduce previously run experiments, pre-load the job submission page
with previously defined configuration, or export a movie of the pseudo-folding process (beta).
A large panel of options is available for running an experiment.
Please refer to the Manual section for a short description of most available options.